Le Infezioni in Medicina, n. 4, 568-573, 2021
doi: 10.53854/liim-2904-10
ORIGINAL ARTICLE
Evaluation of three different methods for susceptibility testing of gentamicin in carbapenem resistant Enterobacterales
Yeliz Tanriverdi Cayci, Kubra Hacieminoglu Ulker, Asuman Birinci
Department of Medical Microbiology, Faculty of Medicine, Ondokuz Mayıs University, Samsun, Turkey
Article received 29 July, 2020; accepted 20 October, 2021
Corresponding author
Yeliz Tanriverdi Cayci
e-mail:yeliztanriverdi@gmail.com
SummaRY
Carbapenem-resistant Enterobacterales (CRE) have become a growing problem worldwide in recent years. Options for the treatment of CRE are limited and one of these options is gentamicin. For this reason, gentamicin susceptibility should be properly determined. In a recently reported study, it is recommended to review the results of automated systems for assessing gentamicin susceptibility in carbapenem-resistant isolates. In this study, we aimed to determine gentamicin susceptibility using three different methods and compare the methods. The study included 107 CRE isolates from different samples. Gentamicin susceptibility was determined using Vitek 2 Compact (bioMérieux, France), Microscan Walkaway Plus (Beckman Coulter, USA) automatic systems, and disk diffusion (DD) method. The broth microdilution method (BMD) was used as reference method. Minor, major, and very major errors and categorical agreement rates were determined for each method. Aminoglycoside-modifying enzymes (aac(6’)Ib and aph(2’’)Ia) were assayed in discrepant isolates. According to BMD results, 90.7%, 1,8 %, and 7.5 % of the isolates were determined as susceptible, intermediate, and resistant to gentamicin, respectively. Compared to the results of the BMD for detecting gentamicin susceptibility, disk diffusion method showed the highest categorical agreement (98.1%), and Vitek 2 Compact showed the lowest categorical agreement (90.6%). The very major error rates were determined 7.5%, 0.9%, and 0.9% for Vitek 2 Compact, Microscan Walkaway Plus, and DD method, respectively. In addition, aac(6’)Ib and aph(2’’)Ia genes were detected in 8 discrepant isolates. For gentamicin susceptibility, the DD showed the most compatible results. The DD can be used as a reliable method for determining gentamicin susceptibility. Compatibility of automated systems with BMD was acceptable, although lower than DD. The discrepancies detected in the Vitek 2 Compact results could be due to the presence of aac(6’)Ib and/or aph(2’’)Ia aminoglycoside-modifying enzymes.
Keywords: Enterobacterales, susceptibility, antimicrobial test, gentamicin, carbapanem resistance.
INTRODUCTION
Enterobacterales family is among the most common pathogens in humans. These bacteria cause syndromes ranging from uncomplicated urinary tract infections to pneumonia, peritonitis, bacteremia, and meningitis. Historically, these bacteria were susceptible to many antibiotics. In recent, however, multiple drug resistance has emerged due to the acquired genes encoding ESBLs, other enzymes, and additional resistance mechanisms [1]. For years, carbapenems have been used as effective agents to treat Enterobacterales infections. However, depending on the use of carbapemes, carbapenem-resistant Enterobacterales (CRE) have emerged in recent years [2].
CRE infections were first reported in the 1990s and are emerging worldwide as a major public health problem [3, 4]. Colistin, fosfomycin, tigecycline, and aminoglycosides are among the limited treatment options for CRE [5-7]. For comparative analysis of antimicrobial agents and successful management of bacterial infections, it is essential to determine the susceptibility to antibiotics correctly [8]. The rapid diagnosis and treatment of microorganisms positively affects patient health and also prevents the emergence of multi-drug resistant strains [9].
Clinical microbiology laboratories use automated or semi-automated systems for the identification and determination of antimicrobial susceptibility [10]. Each system has some strengths and weaknesses; however, the correct identification and determination of susceptibility of the isolates is a fundamental tool for the correct treatment of the patient [11]. These automated systems are based on different analytical methods, such as colorimetry, turbidimetry, or fluorometry, in which bacterial growth is detected on micro-panels containing different substrates for identification and serial dilutions of antibiotics, which in turn allows for establishing minimum inhibitory concentrations (MIC) [12].
The Vitek 2 Compact commercial system uses a turbidimetric method for antimicrobial susceptibility testing. Antimicrobial susceptibility is achieved by means of test cards containing standard dilutions of different antibiotics corresponding to susceptibility cut-off points [12-14]. Susceptibility results can be obtained within 15 hours. On the other hand, Microscan Walkaway Plus panels are 96-well microdilution plates. Susceptibility results can be obtained within 20 hours, depending on the bacteria [9, 14]. The disk diffusion (DD) method is one of the oldest approaches to antimicrobial susceptibility method. It was presented by Kirby Bauer and is also known by this name. This inexpensive and easy-to-apply method is one of the most widely used antimicrobial susceptibility testing methods in routine clinical laboratories [15, 16]. The broth microdilution (BMD) is the standard method used in most reference laboratories [17, 18].
In this study, it was aimed to compare the different methods to determine the susceptibility of gentamicin, which is an alternative antibiotic in the treatment of CRE. The BMD method was used as the reference method.
MATERIALS AND METHODS
Isolates
Enterobacterales isolates, which were isolated from clinical specimens sent to the Ondokuz Mayis University Medical Faculty Hospital Microbiology Laboratory in 2015, and determined carbapenem-resistant with the Vitek 2 Compact (bioMérieux, France) automated system, were included in this study.
Susceptibility testing
Susceptibility testing of gentamicin was performed by Vitek 2 Compact (bioMérieux, France), Microscan Walkaway Plus (Beckman Coulter, USA), and the DD method. The BMD method was used as a reference method.
For Vitek 2 Compact, fresh colonies were used to prepare an inoculum in sterile saline to get 0.45–0.5 McFarland turbidity standard. AST-N325 and AST-N327 antimicrobial susceptibility test cards were used and transferred to the instrument. The cards were automatically read and analyzed by the Vitek computer.
For Microscan Walkaway Plus, fresh colonies were used to prepare an inoculum, and 96 well plates were used. Antibiotic susceptibility was performed in an automated system based on the conventional BMD method.
In the DD method, fresh colonies were used to prepare an inoculum in sterile saline to get 0.45-0.5 McFarland turbidity standard. These bacterial suspensions were inoculated onto a Mueller-Hinton agar plate in three dimensions. Subsequently, an antibiotic disk was placed on the agar. The agar plate containing bacterial inoculum and antibiotic disks were incubated at 35-37°C for 16-24 hours. Antibiotic susceptibility was determined by measuring the diameter of bacterial inhibition zones around the antibiotic disks and interpreted according to the EUCAST criteria [15].
In the BMD method, two-fold dilutions of antibiotics were prepared in a broth growth medium placed in a 96-well microplate. In this method, a cation-adjusted Mueller Hinton broth was used. The antibiotic-containing wells of the microplate were inoculated with a standardized bacterial suspension of 5x105 CFU/ml. After overnight incubation at 35 oC, the microplate was examined for visible bacterial growth. The lowest concentration of antibiotic that prevented growth represented the minimum inhibitory concentration. The results were evaluated according to the EUCAST breakpoints [15]. Finally, all results were compared with the reference method results.
For categorical agreement (CA), very major errors (VME) were defined as those where the test method result was susceptible and the reference method result was resistant, major errors (ME) were defined as those where the test method result was resistant and the reference method result was susceptible, and minor errors (mE) were defined as those where one of the two methods reported a result as intermediate and the other method reported the result as susceptible or resistant. Acceptable performance was evaluated according to criteria of the International Organization for Standardization (ISO) as follows: ≥90% for CA, ≤1.5% for VME, ≤3% for ME, and ≤10% for mE [19]. Escherichia coli ATCC 25922 was used as a quality control for the susceptibility tests.
Molecular analysis
Polymerase chain reaction (PCR) was performed for discrepant isolates that were defined as VME based on BMD results. The genes of aminoglycoside-modifying enzymes aac(6’)Ib and aph(2’’)Ia, which are associated with gentamicin resistance, were investigated by PCR method.
RESULTS
Clinical isolates
A total of 107 Enterobacterales isolated from clinical samples were included in the study. The distribution of the isolated bacteria was as follows: Klebsiella pneumoniae (n=92), Providencia rettgerii (n=6), Escherichia coli (n=3), Enterobacter cloacae (n=2), Klebsiella aerogenes (n=1), Proteus mirabilis (n=1), Morganella morganii (n=1) and Serratia marcescens (n=1).
Susceptibility testing
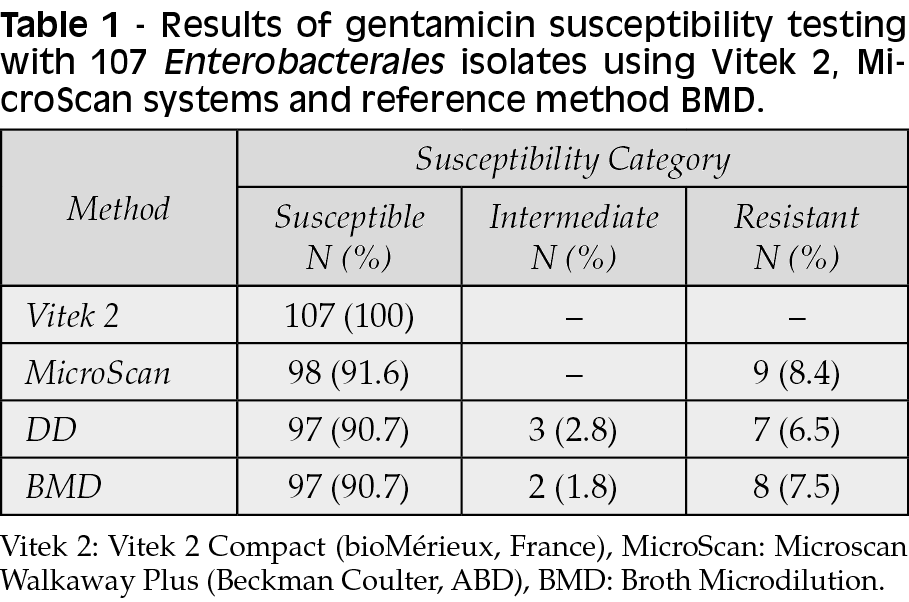
All isolates included in the study were found to be susceptible to gentamicin in the Vitek 2 Compact instrument. Using the Microscan Walkaway Plus instrument, 98 isolates were found to be susceptible to gentamicin and nine isolates were found to be resistant to gentamicin. Using the DD method, 97 isolates were found to be susceptible to gentamicin, seven were found to be resistant to gentamicin, and three isolates were found to be intermediate susceptible to gentamicin. Using the reference method, 97 isolates were found to be susceptible to gentamicin, eight isolates were found to be resistant to gentamicin, and 2 isolates were found to be intermediate susceptible to gentamicin. Discrepant susceptibility results with the reference method were detected in a total of 12 isolates. These results and MIC values are shown in Table 1 and Table 2.
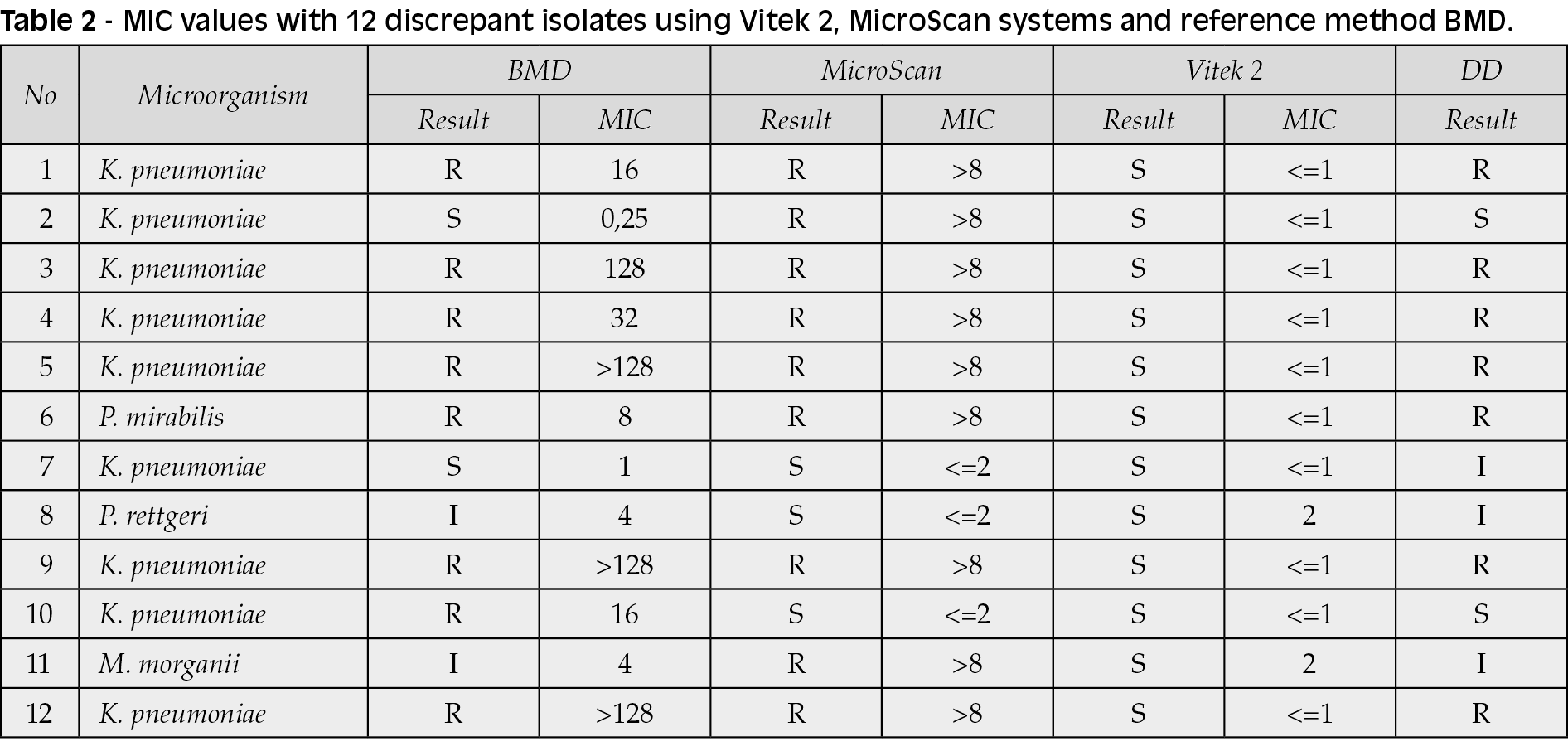
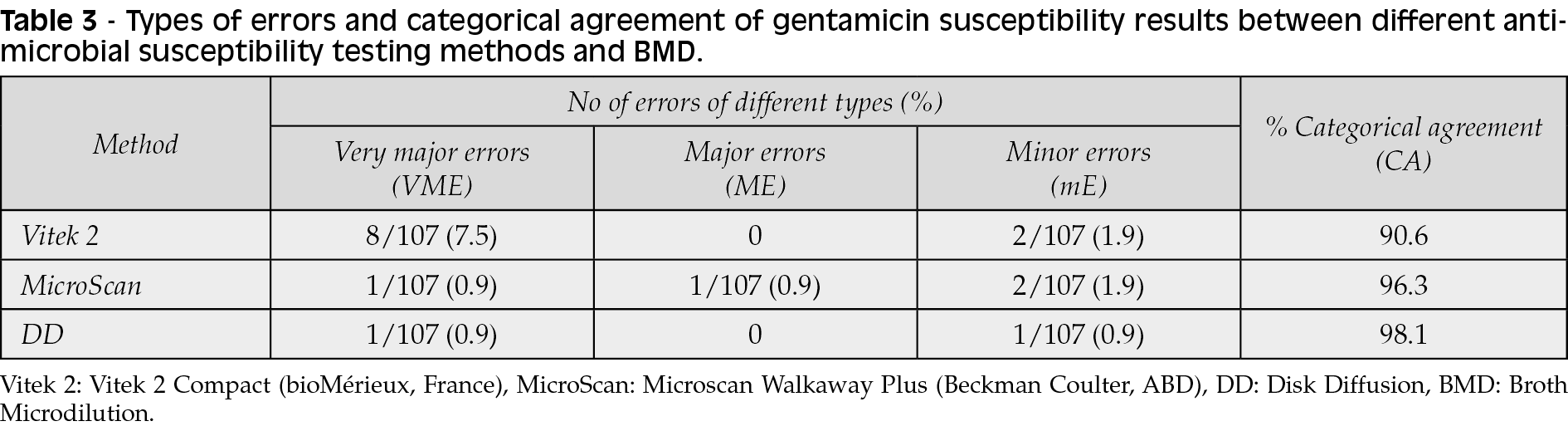
The CA results with a reference method were 90.6%, 96.3%, and 98.1% for Vitek 2 Compact, Microscan Walkaway Plus, and DD method, respectively. No ME was observed with the Vitek 2 Compact system and DD method, while one resistant isolate was found to be susceptible to gentamicin with the MicroScan, resulting in ME (0.9%). Eight resistant isolates were classified as susceptible using Vitek 2, resulting in 8 VMEs (7.5%). One resistant isolate was classified as susceptible using Microscan Walkaway Plus and DD methods, resulting in 1 VME (0.9%) for each method (Table 3).
Molecular analysis
In the Vitek 2 Compact system, VME occurred in 8 isolates. One of these isolates was P. mirabilis, and the others isolates were K. pneumoniae. According to the PCR results of these isolates, all K. pneumoniae isolates had the aminoglycoside-modifying enzymes aac(6’)Ib and aph(2’’)Ia, and the P. mirabilis isolate had the aminoglycoside-modifying enzymes aph(2’’)Ia.
DISCUSSION
Carbapenem-resistant isolates have emerged due to the increased use of carbapenems as a last resort in the treatment of infections caused by these bacteria [1]. Carbapenem resistance is more common in K. pneumoniae isolates than in other Enterobacterales [20]. Gentamicin is a one of the important treatment options for CRE isolates (especially carbapenem-resistant K. pneumoniae) [21]. For this reason, it is important to measure accurately the susceptibility of CRE to gentamicin.
Low accuracy problems for different treatment options have been previously reported in testing antibiotic susceptibility using KPC-KP and other Gram negative bacilli. On the other hand, there are very few studies on the performance of these systems for gentamicin susceptibility testing with similar systems [22-26].
Given the large number of samples of microbiology laboratories today, automated systems such as the Vitek 2 Compact and MicroScan Walkaway Plus have become very popular for antimicrobial susceptibility testing [26]. These commercially available systems, whose main advantages are automated reading and speed, still have lower sensitivity in detecting important resistance mechanisms compared to the DD method [26-31].
In this study, gentamicin susceptibility results showed differences depending on the method used. The DD method showed the best categorical agreement (98.1%) with BMD. On the other hand, Microscan Walkaway Plus and Vitek 2 Compact automated systems showed 96.3% and 90.6% categorical agreement with BMD, respectively. According to the ISO 20776-2 guideline, the performances of the three different methods were considered acceptable [26].
In this study, gentamicin MIC results and categorical agreement results of the Microscan Walkaway Plus showed relatively good agreement with BMD compared to the Vitek 2 Compact. This could be due to differences in the methods of measuring the MICs; the Microscan Walkaway Plus is more similar to the BMD than the Vitek 2 Compact [26]. In our study, the DD method showed the best performance among the reference methods. Excellent agreement was observed between BMD and DD. In previous studies, DD has been validated as a method that shows better agreement with BMD than Vitek 2 Compact and MicroScan. [31].
The discrepancies with associated lower MIC values were found in the Vitek 2 Compact and Microscan Walkaway Plus systems. The MIC values of the VME isolates of the automated systems were lower than the MIC values of the reference method. Similar to our study, isolates reported as VME were found in studies that used EUCAST breakpoints [25, 32]. In another study using CLSI breakpoints, the categorical agreement between Vitek 2 Compact and the reference method was 97% and no isolate was reported as VME [35]. It is known that EUCAST breakpoints for gentamicin are lower than CLSI breakpoints. Since EUCAST breakpoints for susceptibility were used in our study, the discrepancies may have become more apparent.
The eight isolates identified as VME had aac(6’)Ib and aph(2’’)Ia aminoglycoside-modifying enzymes. The Vitek 2 Compact system may have deficiencies in detecting gentamicin resistance associated with these aminoglycoside-modifying enzymes. Previous studies have reported that the Vitek 2 Compact System has deficits in detecting gentamicin resistance due to an armA resistance determinant [25, 34].
Automated methods for susceptibility testing are attractive because of their efficiency, convenience, and a short time to retrieval of results and are widely used in modern clinical microbiology laboratories. In our study, the Vitek 2 Compact and Microscan Walkaway Plus automated systems and the DD method were found to be more than 90% compatible with the reference method. On the other hand, it is thought that the discrepancies in Vitek 2 Compact System may be due to the aac(6’)Ib and/or aph(2’’)Ia aminoglycoside-modifying enzymes and the low EUCAST breakpoints.
Our study has several limitations. The number of isolates tested in this study was relatively small and represented the epidemiology of a single center.
Fundıng
None to declare.
Conflıcts of ınterest
The authors declare that they have no conflicts of interest.
REFERENCES
[1] Schwaber MJ, Carmeli Y. Carbapenem-resistant Enterobacteriaceae: a potential threat. JAMA. 2008; 300 (24), 2911-3.
[2] Morrill HJ, Pogue JM, Kaye KS, LaPlante KL. Treatment options for carbapenem-resistant Enterobacteriaceae infections. Open Forum Infect Dis. 2015; 2 (2).
[3] Gaynes RP, Culver DH. Resistance to imipenem among selected gram-negative bacilli in the United States. Infect Control Hosp Epidemiol. 1992; 13 (1). 10-4.
[4] Nordmann P, Naas T, Poirel L. Global spread of carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis. 2011; 17 (10), 1791-8.
[5] Falagas ME, Lourida P, Poulikakos P, Rafailidis PI, Tansarli GS. Antibiotic treatment of infections due to carbapenem-resistant Enterobacteriaceae: systematic evaluation of the available evidence. Antimicrob Agents Chemother. 2014; 58 (2), 654-63.
[6] Falagas ME, Kasiakou SK. Colistin: the revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin Infect Dis. 2005; 40 (9), 1333-41.
[7] Falagas ME, Kopterides P. Old antibiotics for infections in critically ill patients. Curr Opin Crit Care. 2007; 13 (5), 592-7.
[8] Bonev B, Hooper J, Parisot J. Principles of assessing bacterial susceptibility to antibiotics using the agar diffusion method. J Antimicrob Chemother. 2008; 61 (6), 1295-301.
[9] Jorgensen J.H., Weinstein M.P. Diagnostic technology in clinical microbiology, In Manual Of Clinical Microbiology (Murray PR and Baron EJ., Eds). ASM Press, Washington DC; 2003: 210-11.
[10] Rhoads S, Marinelli L, Imperatrice CA, Nachamkin I. Comparison of Microscan Walkaway Plus system and Vitek system for identifi cation of gram-negative bacteria. J Clin Microbiol. 1995; 33 (11), 3044-6.
[11] U.S. Food and Drug Administration. Guidance for Industry and FDA - Class II Special Controls Guidance Document: Antimicrobial Susceptibility Test (AST) Systems, 2009. Available at: https://www.fda.gov/media/88069/download. [accessed in 5 May 2020].
[12] Hernández-Durán M, López-Jácome LE, Colín-Castro CA, et al. Comparison of the Microscan Walkaway Plus and VITEK 2 Compact systems for the identification and susceptibility of clinical Gram-positive and Gram-negative bacteria. Investigación en Discapacidad. 2017; 6 (3), 105-14.
[13] Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing M100-S27, 2017. Available at: https://clsi.org/media/1469/m100s27_sample.pdf. [accessed in 8 May 2020].
[14] Winstanley T, Courvalin P. Expert systems in clinicalmicrobiology. Clin Microbiol Rev. 2011; 24 (3), 515-56.
[15] The European Committee on Antimicrobial Susceptibility Testing (EUCAST). Breakpoint tables for interpretation of MICs and zone diameters. Version 7.1, 2017. Available at: https://www.eucast.org. [accessed 21 August 2020].
[16] Demirpek U. Antimikrobiyal Duyarlılık Testleri. 2012. Available at: https://www.klimik.org.tr/wp-content/uploads/2012/02/128201112107-49.pdf [accessed in 9 May 2020].
[17] Tenover F.C. Antibiotic susceptibility testing, In Encyclopedia of Microbiology (Schmidt TM., ed). Elsevier Science Publishing Co Inc, San Diego; 2019: 166-75.
[18] Luber P, Bartelt E, Genschow E, Wagner J, Hahn H. Comparison of broth microdilution, E test, and agar dilution methods for antibiotic susceptibility testing of Campylobacter jejuni and Campylobacter coli. J Clin Microbiol. 2003; 41 (3), 1062-8.
[19] International Organization for Standardization. ISO 20776-2. Clinical laboratory testing and in vitro diagnostic test systems. Susceptibility testing of infectious agents and evaluation of performance of antimicrobial susceptibility test devices - Part 2: Evaluation of performance of antimicrobial susceptibility test devices, 2007. [cited in 11 May 2020]. Available from: https://www.iso.org/standard/41631.html
[20] Borah VV, Saikia KK, Chandra P, Hazarika NK, Chakravarty R. New Delhi metallo-b-lactamase and extended spectrum b-lactamases co-producing isolates are high in communityacquired urinary infections in Assam as detected by a novel multiplex polymerasechain reaction assay. Indian J Med Microbiol. 2016; 34 (2), 173-82.
[21] Clancy CJ, Hao B, Shields RK, et al. Doripenem, gentamicin, and colistin, alone and in combinations, against gentamicin-susceptible, KPC-producing Klebsiella pneumoniae strains with various ompK36 genotypes. Antimicrob Agents Chemother. 2014; 58, 3521-5.
[22] Tan TY, Ng SY. Comparison of Etest, Vitek and agar dilution for susceptibility testing of colistin. Clin Microbiol Infect. 2007; 13 (5), 541-4.
[23] Lat A, Clock SA, Wu F, et al. Comparison of polymyxin B, tigecycline, cefepime, and meropenem MICs for KPC-producing Klebsiella pneumoniae by broth microdilution, Vitek 2, and Etest. J Clin Microbiol. 2011; 49 (5), 1795-8.
[24] Zarkotou O, Pournaras S, Altouvas G, et al. Comparative evaluation of tigecycline susceptibility testing methods for expanded-spectrum cephalosporin- and carbapenem-resistant gram-negative pathogens. J Clin Microbiol. 2012; 50 (11), 3747-50.
[25] Arena F, Giani T, Vaggelli G, Terenzi G, Pecile P, Rossolini GM. Accuracy of different methods for susceptibility testing of gentamicin with KPC carbapenemase-producing Klebsiella pneumoniae. Diagn Microbiol Infect Dis. 2015; 81(2): 132-4.
[26] Lee M, Chung HS. Different antimicrobial susceptibility testing methods to detect ertapenem resistance in Enterobacteriaceae: VITEK2, MicroScan, Etest, disk diffusion, and broth microdilution. J Microbiol Methods. 2015; 112, 87-91.
[27] Garrec H, Drieux-Rouzet L, Golmard JL, Jarlier V, Robert J. Comparison of nine phenotypic methods for detection of extended-spectrum beta-lactamase production by Enterobacteriaceae. J Clin Microbiol. 2011; 49, 1048-57.
[28] Lavallee C, Rouleau D, Gaudreau C, et al. Performance of an agar dilution method and a Vitek 2 card for detection of inducible clindamycin resistance in Staphylococcus spp. J Clin Microbiol. 2010; 48 (4), 1354-7.
[29] Tazi A, Réglier-Poupet H, Raymond J, Adam JM, Trieu-Cuot P, Poyart C. Comparative evaluation of VITEK 2 for antimicrobial susceptibility testing of group B Streptococcus. J Antimicrob Chemother. 2007; 59, 1109-13.
[30] Polsfuss S, Bloemberg GV, Giger J, Meyer V, Bottger EC, Hombach M. Practical approach for reliable detection of AmpC beta-Lactamase producing Enterobacteriaceae. J Clin Microbiol. 2011; 49, 2798-803.
[31] Hombach M, Reinhard Z, Böttger EC. Standardisation of disk diffusion results for antibiotic susceptibility testing using the sirscan automated zone reader. BMC Microbiol. 2013; 13 (1), 225.
[32] Ko JH, Baek JY, Peck KR, et al. Discrepant susceptibility to gentamicin despite amikacin resistance in Klebsiella pneumoniae by VITEK 2 represents false susceptibility associated with the armA 16S rRNA methylase gene. J Med Microbiol. 2017; 66, 1448-50.
[33] Calvo J, Cano ME, Pitart C, et al. Evaluation of Three Automated Systems for Susceptibility Testing of Enterobacteria Containing qnrB, qnrS, and/or aac(6’)-Ib-cr. J Clin Microbiol. 2011; 49 (9), 3343-5.
[34] Jung S, Yu JK, Shin SH, et al. False susceptibility to amikacin by VITEK 2 in Acinetobacter baumannii harboring armA. Ann Clin Lab Sci. 2010; 40 (2), 167-71.