Le Infezioni in Medicina, n. 1, 79-84, 2021
ORIGINAL ARTICLES
Rapid identification of non-tuberculous mycobacteria with MALDI-TOF mass spectrometry
Ester Oliva1, Marco Arosio1, Ester Mazzola2, Miriam Mascheroni1, Annarosa Cerro1, Marina Cuntrò1, Francesca Vailati1, Marco Cavallini1, Claudio Farina1
1Microbiology Unit, ASST Papa Giovanni XXIII, Bergamo, Italy;
2Microbiology Unit, ASST Grande Ospedale Metropolitano Niguarda, Milan, Italy
Corresponding author
Claudio Farina
E-mail: cfarina@asst-pg23.it
SummaRY
Rapid and accurate identification of non-tuberculous mycobacteria (NTM) is important for a prompt start to antibiotic therapy. The aim of this study was to obtain accurate identification of NTM quickly by analyzing the performance of the MALDI-TOF mass spectrometry (MS) system VITEK® MS in identifying various NTM species from solid medium and MGIT 960 liquid medium. The study was performed in two phases: preliminary and perspective. Overall, 41/42 species and 33/34 species were correctly identified from the MGIT medium in the preliminary and perspective phases, respectively. The VITEK® MS system includes in its database part of the mycobacteria from the Mycobacterium fortuitum complex but is unable to discriminate among the various species belonging to the complex. Although the VITEK® MS system does not have the protein spectrum of Mycobacterium chimaera, it is not able to distinguish between Mycobacterium chimaera and Mycobacterium intracellulare. Since the VITEK® MS includes the separate protein spectrum of both M. chelonae and M. abscessus, it can discriminate between the two microorganisms. Thanks to these studies we show that the VITEK® MS system is a reliable method for identification of NTMs directly from MGIT liquid medium, instead of the use of solid media.
Keywords: Mycobacteria, MALD-TOF, identification.
INTRODUCTION
The prevalence of Non-Tuberculous Mycobacteria (NTM) infections is growing not only in HIV-positive patients, but also in HIV-negative patients who, however, exhibit a serious lung disease [1, 2].
Whereas Mycobacterium tuberculosis is an obligate pathogen, NTM are environmental organisms which occasionally act as opportunistic pathogens, leading to a wide array of clinical syndromes. However, the Mycobacterium genus comprises more than 150 species. At least 60 NTM species are currently recognized as causative agents of human pathology, with variable severity and prognosis [3, 4].
The official statement of the American Thoracic Society, the Infectious Disease Society of America (ATS/IDSA) and the British Society Thoracic Guidelines on the diagnosis, treatment and prevention of NTM disease specifically recommends identifying - when possible - species-wide strains of clinically significant NTMs [5, 6]. The conventional diagnostic tools for mycobacterial infections - microscopy and culture - remains irreplaceable [7, 8].
New strategies have been developed in the last decades using molecular biology systems based on sequencing or enzymatic restriction targeting the hsp65, 16S rRNA, sod, and rpoB genes: they allow good identification of all mycobacteria at the species level but remain limited to specialized laboratories [9, 10]. In addition, they are expensive, time consuming and require qualified operators. Therefore, diagnostic laboratories should perform species-level identification to establish the clinical significance of a mycobacterial isolate in clinical specimens. Traditional identification of mycobacterial species has mainly relied on phenotypic and biochemical traits [11].
In the last few years, important progress has been made in the application of Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS) for species identification in mycobacteriology. Several features of the Mycobacterium genus, such as the thickness of their cell wall, the lower number of ribosomes and ribosomal proteins compared with other bacteria, or their low growth rate have rendered these microorganisms particularly challenging for MALDI-TOF MS [10]. Unlike most bacteria, mycobacteria require prior inactivation and disruption of the mycolic acid-rich cell wall, both for biosafety reasons and for the release of the cellular protein content.
The BACTEC Mycobacteria Growth Indicator Tube (MGIT) 960 system (Becton Dickinson Cockeysville, Maryland, US) is a fully automated diagnostic system that ensures continuous monitoring to identify positive cultures in real time that has dramatically reduced the time for the etiological diagnosis of mycobacterial infections [13]. In fact, the Bactec MGIT 960 diagnostic system guarantees the rapid growth of mycobacteria both belonging to Mycobacterium tuberculosis complex and to the large group of NTM [14].
The Saramis v4.12 RUO library (bioMérieux, Marcy L’Étoile, France) contains 1286 spectra from 45 Mycobacterium species. Recently, the IVD 3.0 version (VITEK MS, bioMérieux) has also been released, providing robust identification of Mycobacterium isolates both from solid and liquid cultures [14].
The aim of this study was to perform an accurate method of NTM identification using VITEK MS® Mycobacterium/Nocardia Kit (bioMérieux, France), that can reduce timing and the use of expensive molecular methods that are not always available in routine of microbiology laboratories.
To achieve this goal, this study was divided in two phases. First, in the preliminary phase, the method for protein extraction was defined to identify NTM with the MALDI-TOF system (Vitek MS®, bioMérieux). The strains used in this phase were 42 samples present in the collection of the Microbiology and Virology Lab of Bergamo Hospital, 13 of which from the collection of the Lombardy Reference Center for mycobacteria ASST - Grande Ospedale Metropolitano Niguarda, Milan, Italy.
Second, in the perspective phase, the identification of NTM strains was verified directly from the Bactec MGIT 960 liquid medium after the growth without subculture. In this case the samples analyzed were those of patients admitted to ASST-Papa Giovanni XXIII of Bergamo or hospitalized in facilities that refer to the Unit of Microbiology and Virology of Bergamo Hospital for the research of Mycobacterium spp. in biological samples.
The results obtained were compared using mass spectrometry technology (Vitek MS®) proceeding directly from the positive MGIT broths and gene sequencing using 16s rRNA gene, composed of approximately 1,500 nucleotides, which contains most of species specific sequence variations routinely used also for the identification of NTM [9, 10].
MATERIALS AND METHODS
The samples studied in the retrospective phase were 41 clinical isolates of Mycobacterium spp. In particular, the samples below have been studied and identified by gene sequencing using 16s rRNA: Mycobacterium avium (4 strains), Mycobacterium chelonae-abscessus (2), Mycobacterium gordonae (2), Mycobacterium intracellulare (3), Mycobacterium xenopi (3), Mycobacterium fortuitum complex (4), Mycobacterium kansasii (2), Mycobacterium lentiflavum (3), Mycobacterium malmoense (2), Mycobacterium marinum (2), Mycobacterium szulgai (2), Mycobacterium mucogenicum (2), Mycobacterium immunogenum (2), Mycobacterium simiae (1), Mycobacterium goodi (1), Mycobacterium arupense (2), Mycobacterium celatum (1), Mycobacterium scrofulaceum (3). Mycobacterium smegmatis American Type Culture Collection (ATCC) 14468 was used as a control.
For the perspective phase we analyzed the samples directly from culture positive MGIT broths compared with our routine protocol, from April 2018 to September 2018.
All samples were inoculated into the MGIT broth and cultured with MGIT Bactec MGIT 960 instrument as well as International Union Tuberculosis Medium (IUTM, D.I.D. Diagnostic International Distribution, Milan, Italy) and incubated at 35°C. Tubes flagged positive by the MGIT 960 instrument were removed from the instrument, subjected to the acid fast stain to evaluate the growth appearance as purity check and SD Bioline TB Ag MPT4 Rapid test for detection of M. tuberculosis complex.
Two milliliters of the positive MGIT culture broths were transferred to an Eppendorf tube. The tube was centrifuged at 13,000 rpm for 10 minutes to completely remove the supernatant. The pellet was later washed with 500 μL of suspension medium (bioMerieux), centrifuged again before the inactivation. The bacterial pellet was re-suspended with 500 μL of 70% ethanol and transferred to another 1.5 ml Eppendorf tube with 200 μL glass beads (diameter 0.5 mm). Later, it was vortexed for 15 min and kept at room temperature for 10 more minutes. The bacterial suspension was transferred to another 1.5 mL Eppendorf tube, centrifuged for 3 min to completely remove the supernatant. The pellet was mixed with 10 μL 70% formic acid by vortex for 3±5 seconds, then 10 μL 100% acetonitrile was added and mixed again by vortex for 3±5 seconds. The suspension was centrifuged at 13,000 rpm for 2 minutes and 1 μL of the supernatant was spotted onto at VITEK® MS target plate and air dried.
RESULTS
Preliminary phase
As described, the samples were analyzed at the time of growth of the microorganism both on IUTM and on MGIT.
As expected, since the kit by bioMérieux turns out to be validated on IUTM solid medium, the analysis carried out with VITEK® MS allowed to correctly identify 36 NTM at species level and 6 at complex level, out of 42 NTM tested. Starting directly from MGIT, the VITEK® MS identified 35 NTM at species level and 6 at complex level out of 42 of NTM tested, only 1 M. avium was not identified, as described in Table 1.
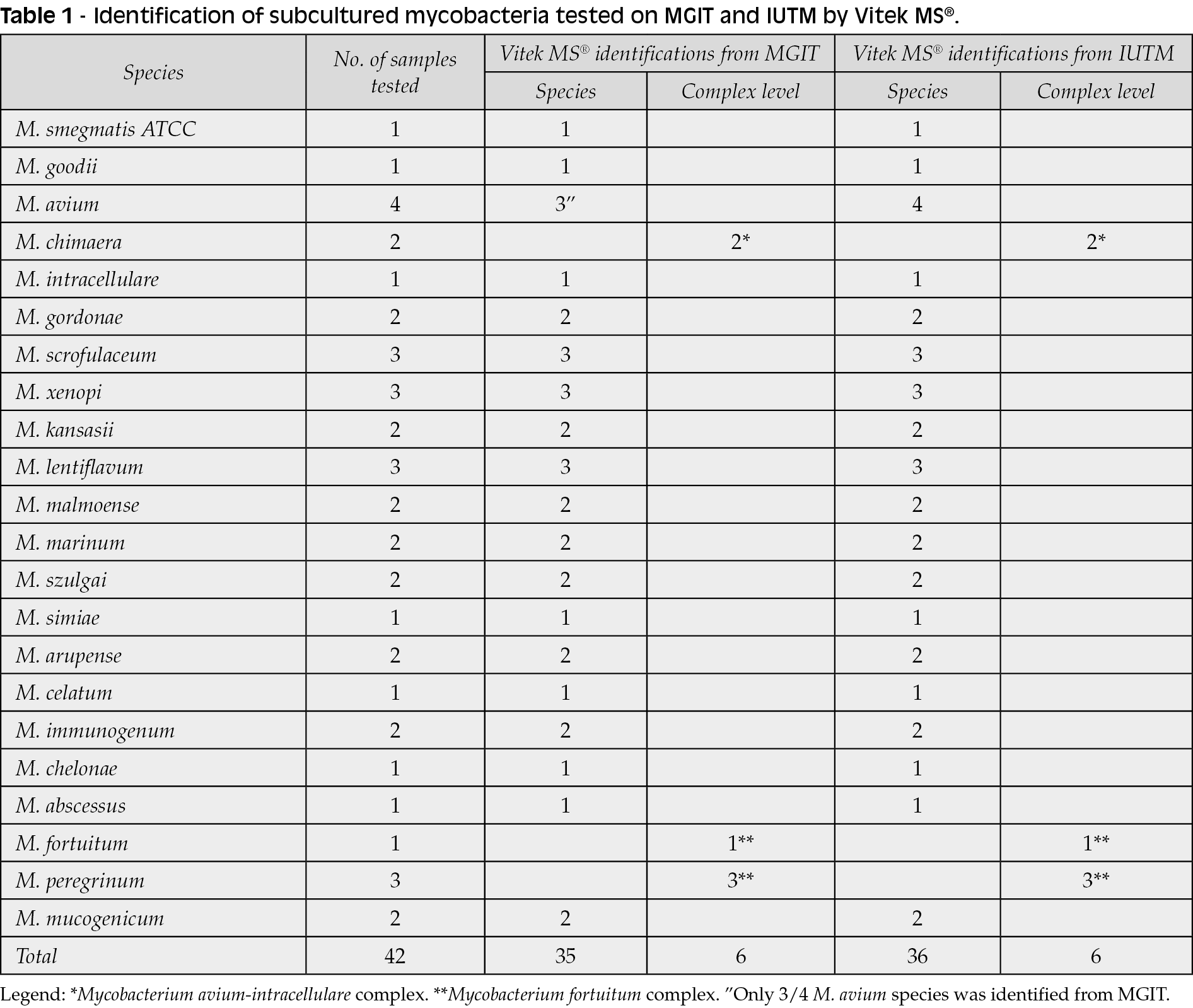
It is important to underline however that, although VITEK® MS system has only included in its RUO (Research Use Only) database most of the mycobacteria belonging to the M. fortuitum complex (M. porcinum, M. peregrinum, M. senegalense, M. houstonense, M. fortuitum ssp fortuitum, M. fortuitum, M. farcinogenes and M. alvei), it is not able to discriminate among the various species belonging to this complex working on Myla™ database (bioMérieux), starting both from MGIT and from IUTM.
Although the VITEK® MS system has not the protein spectrum of Mycobacterium chimaera, is not able to discriminate between Mycobacterium chimaera and Mycobacterium intracellulare both from IUTM and from MGIT.
Perspective phase
We proceeded to identify NTM starting from MGIT all the samples received at the Microbiology and Virology of ASST - Papa Giovanni XXIII, Bergamo for research of Mycobacterium spp.
Every clinical samples have been inoculated, as per protocol, and incubated in MGIT Bactec 960 and in IUTM at 35°C, until the growth. From April 2018 to September 2018, 34 samples were analyzed from 27 different patients. As described in Table 2, VITEK® MS identified at species level 26 out of 34 NTM and 7 out of 34 at complex level.
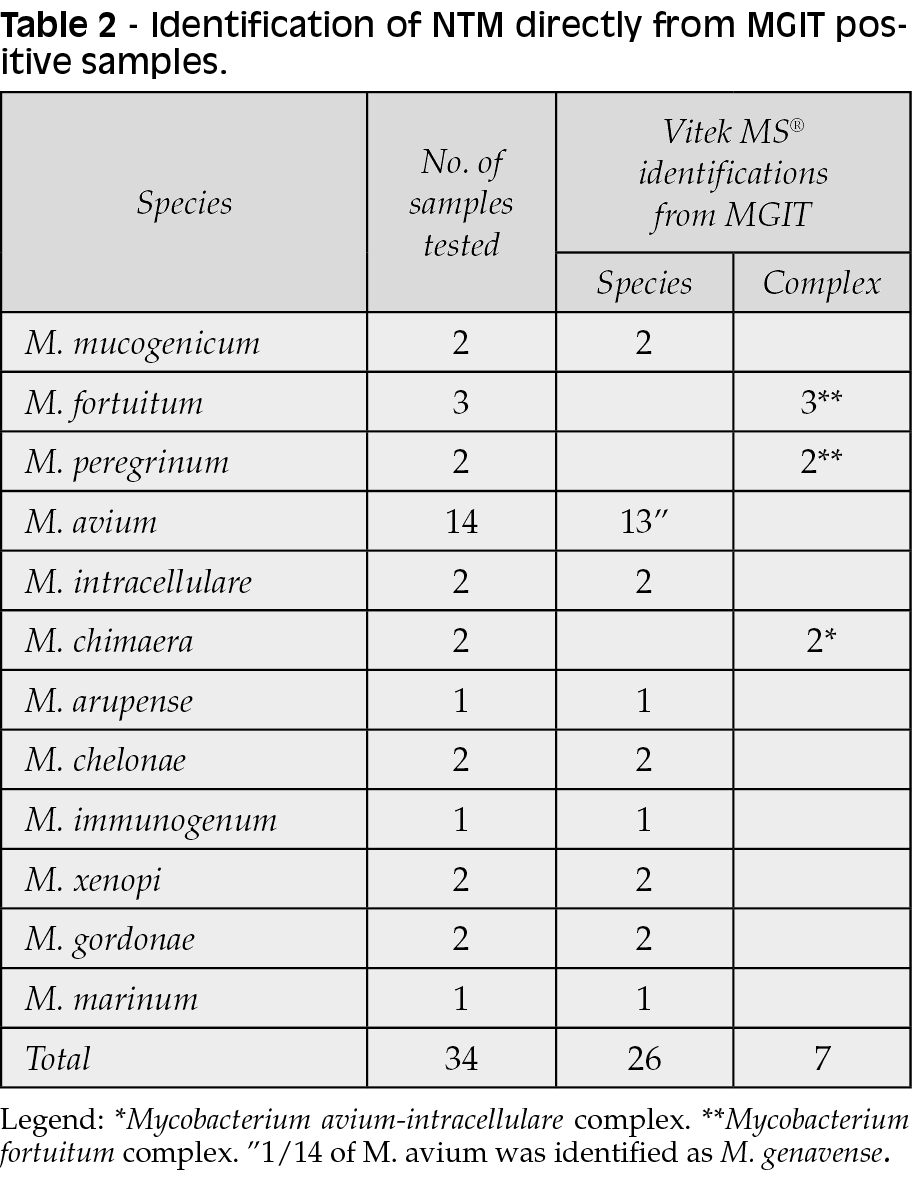
VITEK® MS system did not correctly identify one strain of M. avium isolated from an immunocompromised patient, which was identified as M. genavense. Since the samples are inserted in duplicate, in one of the spots of the VITEK® MS slide, M. genavense was identified at 99.9% and in the other the identification was M. genavense 50%, M. avium 50%. Performing gene sequencing of the 16S rRNA, the microorganism has been identified as M. avium.
Concerning M. intracellulare - chimaera and M. fortuitum complex the same disadvantages that had been detected in the preliminary phase have recurred.
The identifications obtained from the Mycobacterium chelonae-abscessus complex (M. chelonae,
M. immunogenum, M. franklinii, M. salmoniphilum, M. abscessus subsp. abscessus, M. abscessus subsp. massiliense, and M. abscessus subsp. bolletii) appear to be particularly important, as described below, since Vitek® MS system included in its database the separate protein spectra of M. chelonae and M. abscessus, and is thus supposed to be perfectly able to discriminate between the two microorganisms. On the contrary, the gene sequencing of the 16S rRNA is not able to discriminate between the two rapidly growing NTMs.
DISCUSSION
The isolation of NTM from clinical specimens has become very frequent in the last years. Such organisms are typically environmental and poorly pathogenic for humans; they can, however, be responsible for opportunistic diseases in subjects presenting with various predisposing conditions [15, 16]. Timely and accurate identification of NTM species facilitate choices of appropriate antimicrobial therapy [14].
Conventional methods for the identification of NTM were initially classically based on biochemical tests: such technique, however, required several weeks for adequate growth of microorganisms and, sometimes, accurate identification and interpretation were not possible. The growing availability of modern molecular techniques provides microbiologists with extremely powerful tools for accurate identification of microorganisms [11].
Thanks to this study we have shown that Vitek MS® is a reliable method for the identification of NTM at species level directly from positive MGIT broths without previously making subcultures, significantly reducing the identification times of NTM.
As a result of these important improvements, VITEK MS® technology has been implemented for routine identification of NTM in many laboratories and has been shown to perform similarly to conventional methods (DNA amplification-hybridization with species-specific probes) [8-11]. This technique has achieved up to 98% agreement with 16S rRNA, hsp65 and rpoB genes sequencing [8-11].
Due to the presence of separate spectra of M. chelonae and M. abscessus the Vitek MS® system is able to differentiate between the two NTM. Unlike M. chelonae, which in most cases can be considered a contaminant of the sample, M. abscessus must always be considered a pathogen capable of causing infections both in immunocompromised patients and in immunocompetent subjects [16, 17]. Pulmonary infections caused by M. chelonae are rare compared to those by M. abscessus: otherwise, cases of colonization by M. chelonae are known, especially in patients with cystic fibrosis [16].
Furthermore, being able to provide the clinician with information on the identification of M. abscessus in a short time is important, as it would allow a targeted antibiotic therapy to be started as quickly as possible. Therefore, communicating with the clinician and knowing the patient’s concomitant pathologies is important to verify whether the isolated NTM is a contamination or an authentic infection and, in case, request other samples, according with American Thoracic Society criteria and the British Society Thoracic Guidelines [5, 6].
As for the identification of rare microorganisms such as M. genavense - which causes widespread particularly liver, spleen and lymph nodes infections in immunocompromised patients and with complex antibacterial therapies - its initial identification by Vitek MS® has been confirmed, by gene sequencing of the rRNA 16s gene, as M. avium. This misidentification was not found only by our tests, but also in another study, in which M. tuberculosis was identified by the MALDI-TOF system as M. genavense [14]. Since this NTM has very few discriminatory protein peaks [14], we suggest that the identification of this microorganism must always be reconfirmed with validated methods, in such a way as to provide the clinician with the security of a correct identification of species.
The collaboration between the clinician and the microbiologist is crucial. Correct identification is at the basis of a proper treatment which, because of the reduced availability of NTM susceptibility testing, must necessarily be based on data from the literature and consider the different susceptibility patterns that characterize rapidly and slowly growing species [16].
Conflict of interest
Nothing to declare
Funding
None
REFERENCES
[1] Henry MT, Inamdar L, O’Riordain D, Schweiger M, Watson JP. Nontuberculous mycobacteria in non-HIV patients: epidemiology, treatment and response. Eur Resp J. 2004; 23 (5), 741-6.
[2] Lai CC, Tan CK, Chou CH, et al. Increasing Incidence of Nontuberculous Mycobacteria, Taiwan, 2000–2008. Emerg Infect Dis. 2010; 16 (2), 294-6.
[3] Tortoli E. Impact of genotypic studies on mycobacterial taxonomy: the new mycobacteria of the 1990s. Clin Microbiol Rev. 2003; 16 (2), 319-54.
[4] Tortoli E. The new mycobacteria: an update. FEMS Immunology & Medical Microbiology. 2006; 48 (2), 159-78.
[5] Griffith DE, et al. ATS Mycobacterial Diseases Subcommittee; American Thoracic Society; Infectious Disease Society of America. An official ATS/IDSA statement: diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases. Am J Respir Crit Care Med. 2007; 175 (4), 367-416.
[6] Haworth CS, Banks J, Capstick T, et al. British Thoracic Society guidelines for the management of non-tuberculous mycobacterial pulmonary disease (NTM-PD), Thorax. 2017; 72 (Suppl. 2), ii1-ii64.
[7] Salfinger M, Pfyffer GE. The new diagnostic mycobacteriology laboratory. Eur J Clin Microbiol Infect Dis. 1994; 13 (11), 961-79.
[8] Wolinsky E. Conventional Diagnostic Methods for Tuberculosis. Clin Infect Dis. 1994; 19 (3), 396-401.
[9] Brown-Elliott BA, Philley JV. Rapidly growing mycobacteria. Microbiol Spectr. 2017; 5 (1). doi: 10.1128/microbiolspec.TNMI7-0027-2016.
[10] Church DL, Cerutti L, Gürtler A, Griener T, Zelazny A, Emler S. Performance and Application of 16S rRNA gene cycle sequencing for routine identification of bacteria in the clinical microbiology laboratory. Clin Microbiol Rev. 2020; 33 (4), e00053-19.
[11] Lotz A, Ferroni A, Beretti J-L, et al. Rapid identification of mycobacterial whole cells in solid and liquid culture media by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol. 2010; 48 (12), 4481-6.
[12] Alcaide F, Amlerová J, Bou G, et al. How to: identify non-tuberculous Mycobacterium species using MALDI-TOF mass spectrometry. Clin Microbiol Infect. 2018; 24 (6), 599-603.
[13] Huang TS, Chen CS, Lee SS, Huang WK, Liu YC. Comparison of the BACTEC MGIT 960 and BACTEC 460TB systems for detection of mycobacteria in clinical specimens. Ann Clin Lab Science. 2001; 31 (3), 279-83.
[14] Huang TS, Lee CC, Tu HZ, Shin-Jung Lee S. Rapid identification of mycobacteria from positive MGIT broths of primary cultures by MALDI-TOF mass spectrometry. PLoS One. 2018; 13 (2): e0192291.
[15] Tortoli E. Clinical manifestations of nontuberculous mycobacteria infections. Clin Microbiol Infect. 2009; 15 (10), 906-10.
[16] Lee MR, Sheng WH, Hung CC, Yu CJ, Lee LN, Hsueh PR. Mycobacterium abscessus complex infections in humans. Emerg Infect Dis. 2015; 21 (9), 1638-46.
[17] Akram SM, Bhimji SS, Saleh D. Mycobacterium chelonae. Treasure Island (FL): StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2020 Jan. 2019 Jul 3.